Antibodies: a short history
The earliest reference to antibodies dates back to the work of Emil von Behring (who was awarded the Nobel Prize for his work in 1901) and Shibasabura Kitasato in 1890 who showed serum from animals immunized against diphtheria and tetanus to be the cure for infected animals. In 1900 Paul Ehrlich developed his antibody formation theory in agreement the “lock and key” hypothesis that had originally been proposed by Emil Fischer for the interaction of enzymes with their substrates.
The first antibody Isotype (Immunoglobulin G, IgG) was discovered in 1939 by Arne Tiselius and Elvin Kabat and in 1948 Astrid Fagraeus described antibody production in plasma B cells. The molecular structure of antibodies was described 1959 independently by Gerald Edelman and Rodney Porter, for which they shared the Nobel Prize in 1972.
The clonal selection theory was established in 1957 by Frank MacFarlane Burnet and David W. Talmage and extended in 1976 by Susumu Tonegawa’s description of the recombination of Ig genes. 1975 then saw the first generation of hybridomas and development of monoclonal antibodies by Georges Köhler and César Milstein, followed 15 years later by the phage display technique refined by John McCafferty, Andrew D. Griffiths, Greg Winter, and David J Chiswell.
Room for improvement
Antibodies are today a keystone of proteomics research. Antibody production and purification has been refined over the course of the last 125 years. However, antibody production is a resource intensive undertaking.
Immunization of a host animal can be challenging and costly. Highly-toxic or highly-conserved antigens typically elicit an inefficient immune response necessitating use of an adjuvant to spur antibody production. The immunization process takes typically approximately three months. Depending on the immunized animal’s response, additional boost shots might become necessary thus extending the protocol by several weeks until the final bleed.
The fusion of antibody producing B cells isolated from the animal’s spleen and myeloma cells to generate monoclonal antibody producing hybridoma cells adds an additional step of complexity for production of monoclonal antibodies. All of these are independent of subsequent selection, screening and purification steps producing monoclonal antibodies from a newly established hybridoma cell line can easily take a year or more.
Both polyclonal and monoclonal antibody production usually involve sacrificing the immunized animal. Additionally, some hybridomas only grow after having being injected into the peritoneal cavity, secreting a fluid rich in antibodies rich called ascites. This process causes considerable pain to the animal and is therefore avoided if alternative techniques are an option. Consequently, there are ethical issues at play in conventional antibody production.
In addition to production concerns, some of the biochemical characteristics of conventional antibodies also leave room for improvement. Commonly used antibodies like IgG are relatively large with a molecular weight of approximately 150kDa. Much of this weight is incidental, and unnecessary for the antibody’s binding characteristics as the complementary-determining regions (CDR) in the heavy and light chains encompass only a small fractions of the entire antibody. This size becomes an issue in particular for techniques such as immunohistochemistry where antibodies have to pass through barriers like the plasma membrane to reach their designated target.
Finally, the binding affinity of antibodies can vary greatly. Between different lots, variations of several orders of magnitude are common; often from micromolar to picomolar values. Affinity constants of monoclonal antibodies tend to be a bit lower than those of the best binders in population of polyclonal antibodies. On the other hand, polyclonals exhibit higher background signal and show greater lot-to-lot variability.
Alternatives to conventional immunoglobulins
There is mounting pressure to develop and optimize alternatives to traditional immunoglobulins. Both products derived from immunoglobulins and non Ig-based alternatives are rising in popularity, due in large part to the concerns listed above.
Recombinant antibodies
Recombinant antibodies are affinity proteomics reagents that are derived from the sequence of conventional immunoglobulins, but are produced transgenically in a host organism. In contrast to antibodies that are produced by a host animal in response to exposure to an immunogenic substance, recombinant antibodies1 are produced in prokaryotic or eukaryotic expression systems departing from the coding DNA. It is thus assured that the produced antibodies are identical between lots similar to monoclonal antibodies.
Due to the complex heterotetrameric structure of antibodies and the importance of correct post-translational processing such as glycosylation, complete recombinant antibodies are most commonly eukaryotic expression systems. Additional considerations, e.g. reducing conditions present in E. coli cytoplasm, also complicate recombinant antibody production. To some degree, these difficulties have been circumvented by production of smaller antibody fragments that retain the ability to specifically recognize a desired antigen.
Recombinant antibodies and Ig fragments
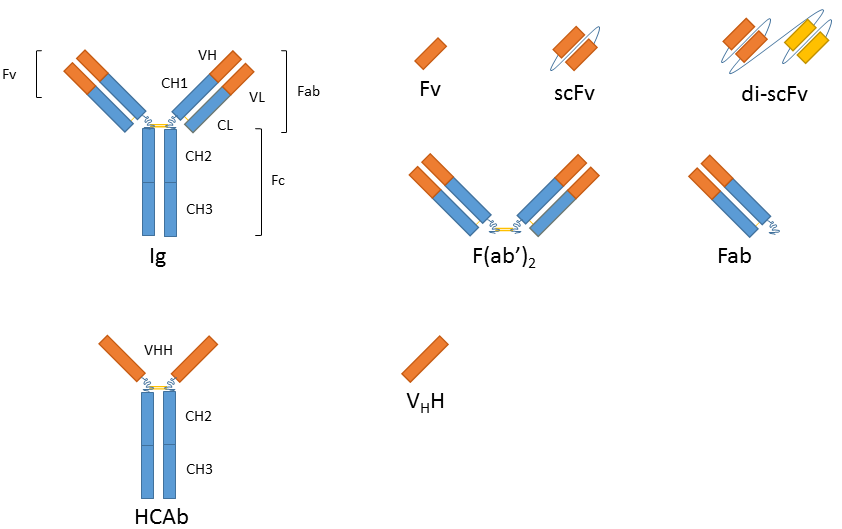
The smallest antigen binding fragment is the immunoglobulin Fv fragment that consists only of one variable domain of the heavy chain. The low stability of the Fv fragment gave rise to the single-chain Fv fragments (scFv) in which one variable region of each light and heavy chain are tethered together by a soluble linker. scFv fragments represent the smallest fragment that holds a complete binding site of an antibody and they are presently the most popular format or recombinant antibodies with wide application in healthcare and biotechnology. Evolution of this platform has also resulted in bi- or tri-valent scFvs and diabodies with lower affinity constants and in bispecific antibodies that can cross-link different antigens and thus hold promise as anti-cancer drugs (e.g. BiTEs). Since all of these variants are produced as a single peptide chain they are still encoded by one gene.
While the absence of the Fc fragment of a full-size antibody can be helpful in certain applications (similar to F(ab’)2 and Fab antibody fragments) it is necessary to retain Ig characteristics in others (e.g. when using a secondary antibody directed against the Fc region). Antibody fragment fusions to the Fc fragment of the immunoglobulin of interest renders the resulting molecule virtually indistinguishable from the native antibody in terms of function and avidity.
Recombinantly produced antibodies have been slowly gathering interest and traction in the research realm. Currently, antibodies-online offers a handful of
against common research targets. Recombinant antibodies against regularly investigated targets like GFP or common loading controls like tubulin make exceptional, regular-use reagents because they display characteristically reproducible results, and have very little lot-to-lot variance with regards to affinity and performance.Furthermore, antibodies-online offers a number of chimeric, partially humanized antibodies against HIV, HCV, and BPV-1 relevant antigens.
In addition to recombinantly produced traditional Ig derivatives, alternatives derived from heavy-chain antibodies (hcAb) are also gaining significant interest. Homodimeric hcAbs of camelids2 and immunoglobulin new antigen receptor (IgNAR)3 of cartilaginous fish consist of only two heavy chains and lack the light chains typical for Ig antibodies. The antigen-binding site of the dromedary HCAb is comprised in one single domain, referred to as variable domain of heavy chain of HCAb (VHH), colloquially referred to as a nanobody4. These nanobodies are particularly small and can be produced using recombinant techniques, with exceptionally high quality and effectively zero lot-to-lot variation.
Nanobodies are small, recombinantly produced antigen binding (VHH) fragments derived from Camelidae. Nanobodies have gathered an incredible following in the research community, and with good reason. The nanobody's unique properties allow it to be utilized in numerous applications, filling roles that conventionally produced antibodies cannot.
Additional Information
Antibody and immunoglobulin alternatives (Part 2) discusses alternatives to conventional mono- and polyclonal antibodies that are not derived from Ig sequences.
References
: "Naturally occurring antibodies devoid of light chains." in: Nature, Vol. 363, Issue 6428, pp. 446-8, (1993) (PubMed).
: "Maturation of shark single-domain (IgNAR) antibodies: evidence for induced-fit binding." in: Journal of molecular biology, Vol. 367, Issue 2, pp. 358-72, (2007) (PubMed).
: "Camelid immunoglobulins and nanobody technology." in: Veterinary immunology and immunopathology, Vol. 128, Issue 1-3, pp. 178-83, (2009) (PubMed).

Goal-oriented, time line driven scientist, proficiently trained in different academic institutions in Germany, France and the USA. Experienced in the life sciences e-commerce environment with a focus on product development and customer relation management.
Go to author page



