The covalent addition of a methyl-group at the 5-carbon of cytosine in DNA, resulting in 5-methylcytosine (5-mC), is an epigenetic mark that influences gene expression, cell differentiation and embryonic development. DNA methyltransferase (DNMT) enzymes catalyze the process. The global methylation pattern in mammals is rather homogenous. However, the CpG islands, which are the main sites for methylation, tend to be near transcription start sites. Furthermore, the methylation level can differ significantly within one organism based on tissue type and between normal and cancerous cell in the same tissue.
While 5-mC is the most common and widely studied DNA-methylation cytosine methylation and demethylation processes give rise to various other forms. In particular 5-hydroxymethylcytosine (5-hmC) has garnered interest since it has been found to be elevated in certain mammalian tissues, such as human and mouse brain, as well as in embryonic stem cells. It is thought to determine developmental fates and, when present in aberrant patterns, contributes to disease. A complete understanding of the potential function 5-hmC requires an accurate, scalable sequencing method.
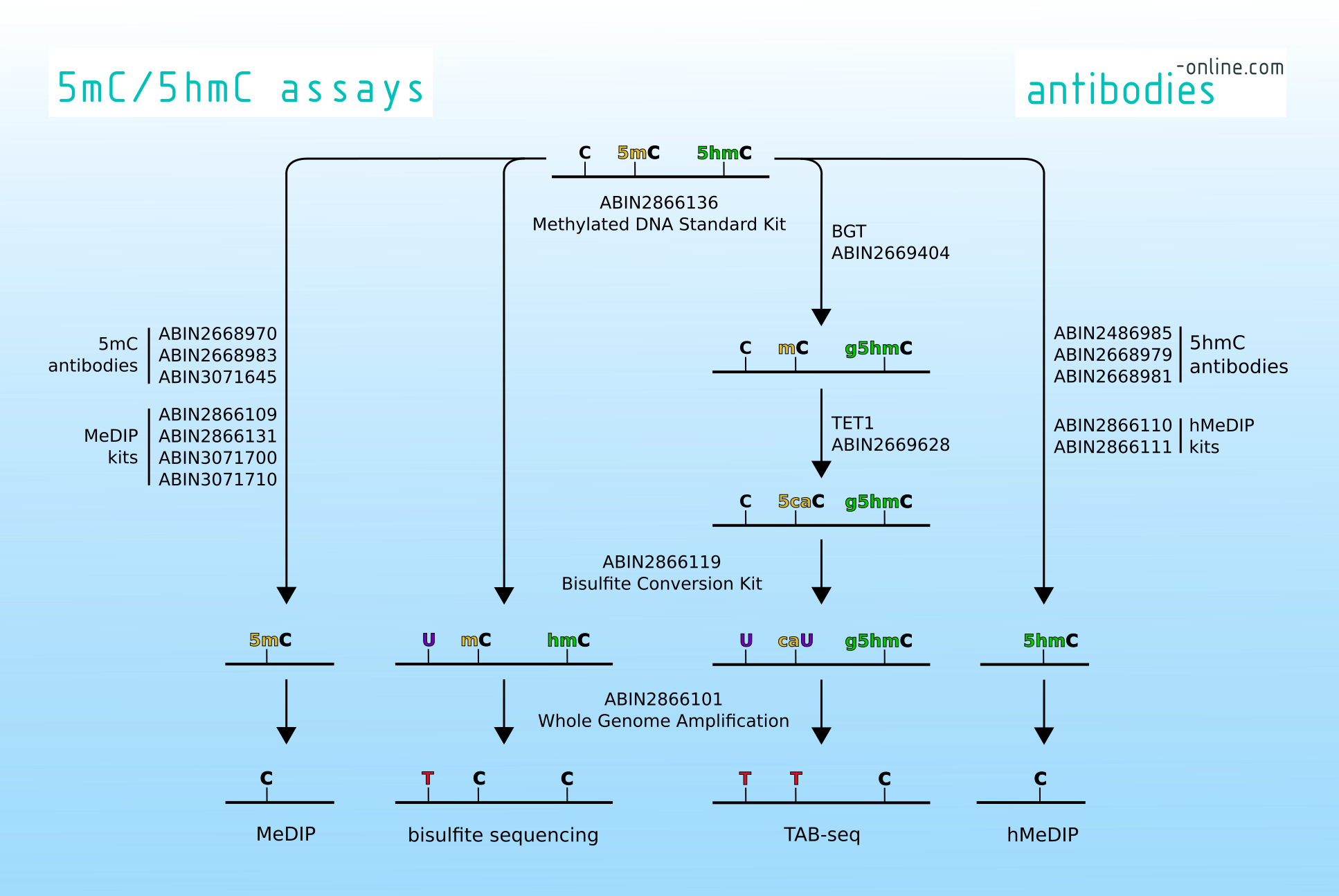
Traditionally, 5-mC sites are identified by bisulfite sequencing or methylated CpG island recovery assay (MIRA). These techniques however cannot distinguish between 5-mC and 5-hmC. TAB-seq (Tet-assisted bisulfite sequencing of 5-hydroxymethylcytosine) is a modification of the standard bisulfite sequencing method through which 5-hydroxymethylcytosine (5-hmC) can be specifically identified. In conjunction with the traditional methods TAB-seq allows mapping of both methylation variants at single-base resolution either on a genomic scale or for a particular locus. TAB-seq was first introduced in 2012. Until now it has been used to study the role of 5-hmC in brain functions, cancers and several cognitive disorders.
Methylated DNA immunoprecipitation (MeDIP) and hydroxylmethylated DNA immunoprecipitation (hMeDIP) on the other hand are two methods that take advantage of 5-mC- and 5-hmC-specific antibodies respectively to enrich DNA fragments bearing these modifications.
antibodies-online is proud to offer products to support your ventures into the DNA methylome with products for MeDIP, hMeDIP, bisulfite sequencing and TAB-seq. Please refer to the table below to find a specific product or get in touch with one of our customer service agents. If you are specifically looking for TAB-seq kits head over to our portal for genomics products genomics-online.
DNA methylation related products
| Product Name | Product Id | Application | Description | Supplier |
| 5-hmC antibody | ABIN2486985 | hMeDIP | hMeDIP validated polyclonal 5-hmC antibody for a successful hMeDIP assay. | Biovision |
| 5-hmC antibody | ABIN3199278 | hMeDIP | hMeDIP validated recombinant 5-hmC antibody for a successful hMeDIP assay. | RevMAb Biosciences |
| 5-mC antibody | ABIN3071645 | MeDIP | MeDIP validated monoclonal 5-mC antibody for a successful MeDIP assay. | Chromatrap |
| 5-mC Antibody with Positive Primer Set | ABIN3071710 | MeDIP | MeDIP validated monoclonal 5-mC antibody and positive primer set provide a complete set of tools for a successful MeDIP assay. | Chromatrap |
| 5-mC Antibody with Positive and Negative Primer Sets | ABIN3071700 | MeDIP | MeDIP validated 5-mC antibody and control primer sets provide a complete set of tools for a successful MeDIP assay. | Chromatrap |
| 5-mC antibody | ABIN2668983 | MeDIP, MeDIP-Seq | Highly characterized MeDIP and MeDIP-Seq validated monoclonal 5-mC antibody. | Active Motif |
| 5-mC antibody | ABIN2668970 | MeDIP | MeDIP validated polyclonal 5-mC antibody for a successful MeDIP assay. | Active Motif |
| 5-hmC antibody | ABIN2668979 | hMeDIP | Highly characterized hMeDIP validated monoclonal 5-hmC antibody. | Active Motif |
| 5-hmC antibody | ABIN2668981 | hMeDIP | Highly characterized hMeDIP validated polyclonal 5-hmC antibody. | Active Motif |
| MeDIP Kit | ABIN2866109 | MeDIP | This MeDIP Kit is a fast, streamlined approach to immunoprecipitate and enrich for single-stranded DNA fragments containing 5-mC. | Active Motif |
| hMeDIP Kit | ABIN2866110 | hMeDIP | This hMeDIP Kit contains a high affinity 5-hmC antibody for efficient immunoprecipitation and enrichment of double or single stranded DNA fragments containing 5-hmC. | Active Motif |
| Hydroxymethyl Collector | ABIN2866111 | hMeDIP | A fast and simple process for enrichment of 5-hmC methylated genomic DNA fragments. | Active Motif |
| HypoMethylCollector | ABIN2866131 | MeDIP, hMeDIP | A novel technology for enrichment of unmethylated genomic DNA fragments for use in many downstream applications. | Active Motif |
| MethylCollector Ultra | ABIN2866132 | MeDIP | A novel technology capable of quickly and efficiently isolating methylated CpG islands from fragmented genomic DNA. | Active Motif |
| β-Glucosyltransferase | ABIN2669404 | TAB-seq | β-Glucosyltransferase can be used to glucosylate 5-hmC DNA. | Active Motif |
| Tet Methylcytosine Dioxygenase 1 (TET1) | ABIN2669628 | TAB-seq | Tet1 is a cytosine oxygenase that catalyzes the conversion of the modified genomic base 5-mC into 5-hydroxymethylcytosine (5-hmC). | Active Motif |
| Methylated DNA Standard Kit | ABIN2866136 | MeDIP, hMeDIP | DNA standards that can be used as positive controls in methylation analysis experiments. | Active Motif |
| Bisulfite Conversion Kit | ABIN2866119 | Bisulfite sequencing, TAB-seq | Bisulfite Conversion Kit that simplifies the analysis of DNA methylation by providing optimized conversion reagents and an easy-to-use protocol. | Active Motif |
| GenoMatrix Whole Genome Amplification Kit | ABIN2866101 | Whole Genome Amplification | Bias-free amplification of human genomic DNA. | Active Motif |
References
: "Base-resolution analysis of 5-hydroxymethylcytosine in the mammalian genome." in: Cell, Vol. 149, Issue 6, pp. 1368-80, (2012) (PubMed).
: "Tet-assisted bisulfite sequencing of 5-hydroxymethylcytosine." in: Nature protocols, Vol. 7, Issue 12, pp. 2159-70, (2013) (PubMed).
: "DNA methylation and demethylation in mammals." in: The Journal of biological chemistry, Vol. 286, Issue 21, pp. 18347-53, (2011) (PubMed).

Goal-oriented, time line driven scientist, proficiently trained in different academic institutions in Germany, France and the USA. Experienced in the life sciences e-commerce environment with a focus on product development and customer relation management.
Go to author page